Genetic code
Background to the schools Wikipedia
Arranging a Wikipedia selection for schools in the developing world without internet was an initiative by SOS Children. SOS Child sponsorship is cool!
The genetic code is the set of rules by which information encoded in genetic material (DNA or RNA sequences) is translated into proteins (amino acid sequences) by living cells. Specifically, the code defines a mapping between tri- nucleotide sequences called codons, and amino acids; every triplet of nucleotides in a nucleic acid sequence specifies a single amino acid. Because the vast majority of genes are encoded with exactly the same code (see #RNA codon table), this particular code is often referred to as the canonical or standard genetic code, or simply the genetic code, though in fact there are many variant codes; thus, the canonical genetic code is not universal. For example, in humans, protein synthesis in mitochondria relies on a genetic code that varies from the canonical code.
It is important to know that not all genetic information is stored as the genetic code. All organisms' DNA contain regulatory sequences, intergenic segments, chromosomal structural areas, which can contribute greatly to phenotype but operate using a distinct sets of rules which may or may not be as straightforward as the well-defined codon-to-amino acid paradigm which underlies the genetic code.
Cracking the genetic code
After the structure of DNA was deciphered by James Watson, Francis Crick, Maurice Wilkins and Rosalind Franklin, serious efforts to understand the nature of the encoding of proteins began. George Gamov postulated that a three-letter code must be employed to encode the 20 different amino acids used by living cells to encode proteins (because 3 is the smallest n such that 4n is at least 20). The fact that codons did consist of three DNA bases was first demonstrated in the Crick, Brenner et al. experiment. The first elucidation of a codon was done by Marshall Nirenberg and Heinrich J. Matthaei in 1961 at the National Institutes of Health. They used a cell-free system to translate a poly-uracil RNA sequence (or UUUUU... in biochemical terms) and discovered that the polypeptide they had synthesized consisted of only the amino acid phenylalanine. They thereby deduced from this poly-phenylalanine that the codon UUU specified the amino-acid phenylalanine. Extending this work, Nirenberg and his coworkers were able to determine the nucleotide makeup of each codon. In order to determine the order of the sequence, trinucleotides were bound to ribosomes and radioactively labeled aminoacyl-tRNA was used to determine which amino acid corresponded to the codon. Nirenberg's group was able to determine the sequences of 54 out of 64 codons. Subsequent work by Har Gobind Khorana identified the rest of the code, and shortly thereafter Robert W. Holley determined the structure of transfer RNA, the adapter molecule that facilitates translation. This work was based upon earlier studies by Severo Ochoa, who received the Nobel prize in 1959 for his work on the enzymology of RNA synthesis. In 1968, Khorana, Holley and Nirenberg also received the Nobel Prize in Physiology or Medicine for their work.
Transfer of information via the genetic code
The genome of an organism is inscribed in DNA, or in some viruses RNA. The portion of the genome that codes for a protein or an RNA is referred to as a gene. Those genes that code for proteins are composed of tri-nucleotide units called codons, each coding for a single amino acid. Each nucleotide sub-unit consists of a phosphate, deoxyribose sugar and one of the 4 nitrogenous nucleotide bases. The purine bases adenine (A) and guanine (G) are larger and consist of two aromatic rings. The pyrimidine bases cytosine (C) and thymine (T) are smaller and consist of only one aromatic ring. In the double-helix configuration, two strands of DNA are joined to each other by hydrogen bonds in an arrangement known as base pairing. These bonds almost always form between an adenine base on one strand and a thymine on the other strand and between a cytosine base on one strand and a guanine base on the other. This means that the number of A and T residues will be the same in a given double helix as will the number of G and C residues. In RNA, thymine (T) is replaced by uracil (U), and the deoxyribose is substituted by ribose.
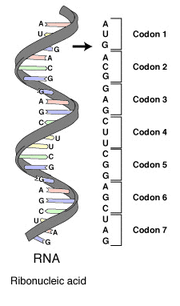
Each protein-coding gene is transcribed into a template molecule of the related polymer RNA, known as messenger RNA or mRNA. This in turn is translated on the ribosome into an amino acid chain or polypeptide. The process of translation requires transfer RNAs specific for individual amino acids with the amino acids covalently attached to them, guanosine triphosphate as an energy source, and a number of translation factors. tRNAs have anticodons complementary to the codons in mRNA and can be "charged" covalently with amino acids at their 3' terminal CCA ends. Individual tRNAs are charged with specific amino acids by enzymes known as aminoacyl tRNA synthetases which have high specificity for both their cognate amino acids and tRNAs. The high specificity of these enzymes is a major reason why the fidelity of protein translation is maintained.
There are 4³ = 64 different codon combinations possible with a triplet codon of three nucleotides. In reality, all 64 codons of the standard genetic code are assigned for either amino acids or stop signals during translation. If, for example, an RNA sequence, UUUAAACCC is considered and the reading-frame starts with the first U (by convention, 5' to 3'), there are three codons, namely, UUU, AAA and CCC, each of which specifies one amino acid. This RNA sequence will be translated into an amino acid sequence, three amino acids long. A comparison may be made with computer science, where the codon is the equivalent of a word, which is the standard "chunk" for handling data (like one amino acid of a protein), and a nucleotide for a bit.
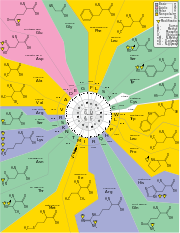
The standard genetic code is shown in the following tables. Table 1 shows what amino acid each of the 64 codons specifies. Table 2 shows what codons specify each of the 20 standard amino acids involved in translation. These are called forward and reverse codon tables, respectively. For example, the codon AAU represents the amino acid asparagine, and UGU and UGC represent cysteine (standard three-letter designations, Asn and Cys respectively).
RNA codon table
| 2nd base | |||||
|---|---|---|---|---|---|
| U | C | A | G | ||
| 1st base |
U |
UUU (Phe/F) Phenylalanine |
UCU (Ser/S) Serine |
UAU (Tyr/Y) Tyrosine |
UGU (Cys/C) Cysteine |
| C |
CUU (Leu/L)Leucine |
CCU (Pro/P) Proline |
CAU (His/H) Histidine |
CGU (Arg/R) Arginine |
|
| A |
AUU (Ile/I) Isoleucine |
ACU (Thr/T) Threonine |
AAU (Asn/N) Asparagine |
AGU (Ser/S)Serine |
|
| G |
GUU (Val/V) Valine |
GCU (Ala/A) Alanine |
GAU (Asp/D) Aspartic acid |
GGU (Gly/G) Glycine |
|
| Ala/A | GCU, GCC, GCA, GCG | Leu/L | UUA, UUG, CUU, CUC, CUA, CUG |
|---|---|---|---|
| Arg/R | CGU, CGC, CGA, CGG, AGA, AGG | Lys/K | AAA, AAG |
| Asn/N | AAU, AAC | Met/M | AUG |
| Asp/D | GAU, GAC | Phe/F | UUU, UUC |
| Cys/C | UGU, UGC | Pro/P | CCU, CCC, CCA, CCG |
| Gln/Q | CAA, CAG | Ser/S | UCU, UCC, UCA, UCG, AGU, AGC |
| Glu/E | GAA, GAG | Thr/T | ACU, ACC, ACA, ACG |
| Gly/G | GGU, GGC, GGA, GGG | Trp/W | UGG |
| His/H | CAU, CAC | Tyr/Y | UAU, UAC |
| Ile/I | AUU, AUC, AUA | Val/V | GUU, GUC, GUA, GUG |
| START | AUG | STOP | UAG, UGA, UAA |
Salient features
Reading frame of a sequence
Note that a codon is defined by the initial nucleotide from which translation starts. For example, the string GGGAAACCC, if read from the first position, contains the codons GGG, AAA and CCC; and if read from the second position, it contains the codons GGA and AAC; if read starting from the third position, GAA and ACC. Partial codons have been ignored in this example. Every sequence can thus be read in three reading frames, each of which will produce a different amino acid sequence (in the given example, Gly-Lys-Pro, Gly-Asp, or Glu-Thr, respectively). With double-stranded DNA there are six possible reading frames, three in the forward orientation on one strand and three reverse (on the opposite strand).
The actual frame in which a protein sequence is translated is defined by a start codon, usually the first AUG codon in the mRNA sequence. Mutations that disrupt the reading frame by insertions or deletions of a non-multiple of 3 nucleotide bases are known as frameshift mutations. These mutations may impair the function of the resulting protein, if it is formed, and are thus rare in in vivo protein-coding sequences. Often such misformed proteins are targeted for proteolytic degradation. In addition, a frame shift mutation is very likely to cause a stop codon to be read which truncates the creation of the protein (example ). One reason for the rareness of frame-shifted mutations being inherited is that if the protein being translated is essential for growth under the selective pressures the organism faces, absence of a functional protein may cause lethality before the organism is viable.
Start/stop codons
Translation starts with a chain initiation codon (start codon). Unlike stop codons, the codon alone is not sufficient to begin the process. Nearby sequences and initiation factors are also required to start translation. The most common start codon is AUG, which codes for methionine, so most amino acid chains start with methionine.
The three stop codons have been given names: UAG is amber, UGA is opal (sometimes also called umber), and UAA is ochre. "Amber" was named by discoverers Richard Epstein and Charles Steinberg after their friend Harris Bernstein, whose last name means "amber" in German. The other two stop codons were named 'ochre" and "opal" in order to keep the "colour names" theme. Stop codons are also called termination codons and they signal release of the nascent polypeptide from the ribosome due to binding of release factors in the absence of cognate tRNAs with anticodons complementary to these stop signals.
Degeneracy of the genetic code
The genetic code has redundancy but no ambiguity (see the codon tables above for the full correlation). For example, although codons GAA and GAG both specify glutamic acid (redundancy), neither of them specifies any other amino acid (no ambiguity). The codons encoding one amino acid may differ in any of their three positions. For example the amino acid glutamic acid is specified by GAA and GAG codons (difference in the third position), the amino acid leucine is specified by UUA, UUG, CUU, CUC, CUA, CUG codons (difference in the first or third position), while the amino acid serine is specified by UCA, UCG, UCC, UCU, AGU, AGC (difference in the first, second or third position).
A position of a codon is said to be a fourfold degenerate site if any nucleotide at this position specifies the same amino acid. For example, the third position of the glycine codons (GGA, GGG, GGC, GGU) is a fourfold degenerate site, because all nucleotide substitutions at this site are synonymous, i.e. they do not change the amino acid. Only the third positions of some codons may be fourfold degenerate. A position of a codon is said to be a twofold degenerate site if only two of four possible nucleotides at this position specify the same amino acid. For example, the third position of the glutamic acid codons (GAA, GAG) is a twofold degenerate site, so is the first position of the leucine codons (UCA, UCC, CCU, CCC, CCA, CCG). In twofold degenerate sites, the equivalent nucleotides are always either two purines (A/G) or two pyrimidines (C/U), so only transversional substitutions (purine to pyrimidine or pyrimidine to purine) in twofold degenerate sites are nonsynonymous. A position of a codon is said to be a non-degenerate site if any mutation at this position results in amino acid substitution. There is only one threefold degenerate site where changing three of the four nucleotides has no effect on the amino acid, while changing the fourth possible nucleotide results in an amino acid substitution. This is the third position of an isoleucine codon: AUU, AUC, or AUA all encode isoleucine, but AUG encodes methionine. In computation this position is often treated as a twofold degenerate site.
There are three amino acids encoded by six different codons: serine, leucine, arginine. Only two amino acids are specified by a single codon; one of these is the amino-acid methionine, specified by the codon AUG, which also specifies the start of translation; the other is tryptophan, specified by the codon UGG. The degeneracy of the genetic code is what accounts for the existence of silent mutations.
Degeneracy results because a triplet code designates 20 amino acids and a stop codon. Because there are four bases, triplet codons are required to produce at least 21 different codes. For example, if there were two bases per codon, then only 16 amino acids could be coded for (4²=16). Because at least 21 codes are required, then 4³ gives 64 possible codons, meaning that some degeneracy must exist.
These properties of the genetic code make it more fault-tolerant for point mutations. For example, in theory, fourfold degenerate codons can tolerate any point mutation at the third position, although codon usage bias restricts this in practice in many organisms; twofold degenerate codons can tolerate one out of the three possible point mutations at the third position. Since transition mutations (purine to purine or pyrimidine to pyrimidine mutations) are more likely than transversion (purine to pyrimidine or vice-versa) mutations, the equivalence of purines or that of pyrimidines at twofold degenerate sites adds a further fault-tolerance.
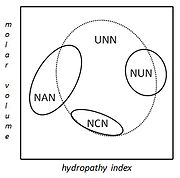
A practical consequence of redundancy is that some errors in the genetic code only cause a silent mutation or an error that would not affect the protein because the hydrophilicity or hydrophobicity is maintained by equivalent substitution of amino acids; for example, a codon of NUN (where N = any nucleotide) tends to code for hydrophobic amino acids. NCN yields amino acid residues that are small in size and moderate in hydropathy; NAN encodes average size hydrophilic residues; UNN encodes residues that are not hydrophilic.
Even so, single point mutations can still cause dysfunctional proteins. For example, a mutated hemoglobin gene causes sickle-cell disease. In the mutant hemoglobin a hydrophilic glutamate (Glu) is substituted by the hydrophobic valine (Val), which reduces the solubility of β-globin. In this case, this mutation causes hemoglobin to form linear polymers linked by the hydrophobic interaction between the valine groups causing sickle-cell deformation of erythrocytes. Sickle-cell disease is generally not caused by a de novo mutation. Rather it is selected for in malarial regions (in a way similar to thalassemia), as heterozygous people have some resistance to the malarial Plasmodium parasite ( heterozygote advantage).
These variable codes for amino acids are allowed because of modified bases in the first base of the anticodon of the tRNA, and the base-pair formed is called a wobble base pair. The modified bases include inosine and the Non-Watson-Crick U-G basepair.
Variations to the standard genetic code
While slight variations on the standard code had been predicted earlier, none were discovered until 1979, when researchers studying human mitochondrial genes discovered they used an alternative code. Many slight variants have been discovered since, including various alternative mitochondrial codes, as well as small variants such as Mycoplasma translating the codon UGA as tryptophan. In bacteria and archaea, GUG and UUG are common start codons. However, in rare cases, certain specific proteins may use alternative initiation (start) codons not normally used by that species.
In certain proteins, non-standard amino acids are substituted for standard stop codons, depending upon associated signal sequences in the messenger RNA: UGA can code for selenocysteine and UAG can code for pyrrolysine as discussed in the relevant articles. Selenocysteine is now viewed as the 21st amino acid, and pyrrolysine is viewed as the 22nd. A detailed description of variations in the genetic code can be found at the NCBI web site.
Notwithstanding these differences, all known codes have strong similarities to each other, and the coding mechanism is the same for all organisms: three-base codons, tRNA, ribosomes, reading the code in the same direction and translating the code three letters at a time into sequences of amino acids.
Theories on the origin of the genetic code
Despite the variations that exist, the genetic codes used by all known forms of life on Earth are very similar. Since there are many possible genetic codes that are thought to have similar utility to the one used by Earth life, the theory of evolution suggests that the genetic code was established very early in the history of life, with phylogenetic analysis of transfer RNA suggests that tRNA molecules evolved before the present set of aminoacyl-tRNA synthetases.
The genetic code is not a random assignment of codons to amino acids. For example, amino acids that share the same biosynthetic pathway tend to have the same first base in their codons, and amino acids with similar physical properties tend to have similar codons.
There are three themes running through the many theories that seek to explain the evolution of the genetic code (and hence the origin of these patterns). One is illustrated by recent aptamer experiments which show that some amino acids have a selective chemical affinity for the base triplets that code for them. This suggests that the current, complex translation mechanism involving tRNA and associated enzymes may be a later development, and that originally, protein sequences were directly templated on base sequences. Another is that the standard genetic code that we see today grew from a simpler, earlier code through a process of "biosynthetic expansion". Here the idea is that primordial life 'discovered' new amino acids (e.g. as by-products of metabolism) and later back-incorporated some of these into the machinery of genetic coding. Although much circumstantial evidence has been found to suggest that fewer different amino acids were used in the past than today, precise and detailed hypotheses about exactly which amino acids entered the code in exactly what order has proved far more controversial. A third theory is that natural selection has led to codon assignments of the genetic code that minimize the effects of mutations.